In-Fusion® HD Cloning Kit
使用文献
In-Fusion Cloningシリーズ関連の使用文献集
タカラバイオのIn-Fusionクローニングシステムは、どんなベクターのどんな位置にもディレクショナルクローニングが可能で、クローニング効率が高く、質の高いクローニングを行うことができるシステムです。下記をはじめとする様々な分野で実績があり、In-Fusionクローニングシステムを使用した文献は年々増え続けています。- In-Fusionを用いてプラスミドベクターを構築した例
- CRISPR/Cas9にIn-Fusionが用いられた例
- ハイスループットスクリーニングにIn-Fusionが用いられた例
- 次世代シーケンシング(NGS)にIn-Fusionが用いられた例
● In-Fusion Cloningシリーズ 使用文献数*
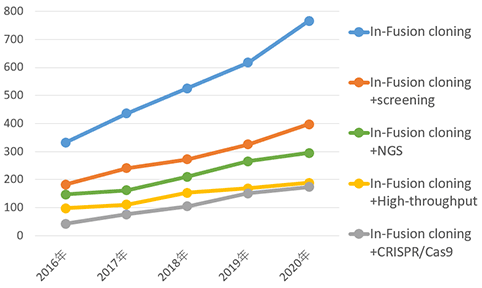
* 論文数はGoogle Scholarにより集計した。

* 論文数はGoogle Scholarにより集計した。
1、In-Fusionを用いてプラスミドベクターを構築した例
- Construction and characterization of centromeric plasmids for Komagataella phaffii using a color-based plasmid stability assay.
Piva LC., et al. PLoS One. (2020) 15(7): e0235532. - Cloning short DNA into plasmids by one-step PCR.
Tao CC., et al. Thoracic Cancer. (2020) 11(11):3409-3415. - A standard for near-scarless plasmid construction using reusable DNA parts.
Ma X., et al. Nat Commun. (2019) 10:3294. - A novel, easy and rapid method for constructing yeast two-hybrid vectors using In-Fusion technology.
Yu D., et al. Biotechniques. (2018) 64(5):219-224. - Optimizing the DNA Donor Template for Homology-Directed Repair of Double-Strand Breaks.
Fei Song & Knut Stieger, Nucleic Acids. (2017) 7:53-60.
2、CRISPR/Cas9にIn-Fusionが用いられた例
- Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping.
Gee P., et al. Nat Commun. (2020) 11(1):1334. - Functional analysis and development of a CRISPR/Cas9 allelic series for a CPR5 ortholog necessary for proper growth of soybean trichomes.
Campbell B.W., et al. Sci Rep. (2019) 9:14757. - Efficient CRISPR/Cas9-based genome editing and its application to conditional genetic analysis in Marchantia polymorpha.
Shigeno S., et al. PLoS One. (2018)13(10): e0205117. - A highly efficient ligation-independent cloning system for CRISPR/Cas9 based genome editing in plants.
Khan, A.A., et al. Plant Methods. (2017) 13,:86. - Efficient genomic correction methods in human iPS cells using CRISPR–Cas9 system.
Li H.L., et al. Methods. (2016) 101:27-35. - MMEJ-assisted gene knock-in using TALENs and CRISPR-Cas9 with the PITCh systems.
Sakuma T., et al. Nat Protoc. (2016) 11(1):118-133. - Targeted genome modifications in soybean with CRISPR/Cas9.
Jacobs, et al. BMC Biotechnology. (2015) 15:16.
3、ハイスループットスクリーニングにIn-Fusionが用いられた例
- Rapid and efficient enhancer cloning and in vivo screening using the developing chick embryo.
Ruth M. Williams & Tatjana Sauka-Spengler, STAR Ptotoc. (2021) 2(2):100507. - High throughput instrument to screen fluorescent proteins under two-photon excitation.
ROSANA S. et al. Biomed Opt Express. (2020) 11(12):7192-7203. - Construction of gateway-compatible baculovirus expression vectors for high-throughput protein expression and in vivo microcrystal screening.
Tang Y., et al. Sci Rep. (2020) 10: 13323. - Serodiagnostic antigens of Clonorchis sinensis identified and evaluated by high-throughput proteogenomics.
Cho PY., et al. PLoS Negl Trop Dis. (2020) 14(12): e0008998. - High throughput generation and characterization of replication-competent clade C transmitter-founder simian human immunodeficiency viruses.
Dutta D., et al. PLoS One. (2018) 13(5):e0196942. - Rapid high-throughput cloning and stable expression of antibodies in HEK293 cells.
Spidel J.L., et al. J Immunol Methods. (2016) 439:50-58. - A versatile ligation-independent cloning method suitable for high-throughput expression screening applications.
Berrow N.S., et al. Nucleic Acids Res. (2007) 35(6):e45. - Many paths to many clones: a comparative look at high-throughput cloning methods.
Marsischky G. & LaBaer J., Genome Res. (2004) 14:2020-2028.
4、次世代シーケンシング(NGS)にIn-Fusionが用いられた例
- Preclinical assessment of the efficacy and specificity of GD2-B7H3 SynNotch CAR-T in metastatic neuroblastoma.
Moghimi B., et al. Nat Commun. (2021) 12:511. - An automated pipeline for the screening of diverse monoterpene synthase libraries.
Leferink, et al. Sci Rep. (2019) 9.:11936. - Strategy for efficient generation of numerous full-length cDNA clones of classical swine fever virus for haplotyping.
Johnston, CM., et al. BMC Genomics. (2018) 19(1)600.
